Guide
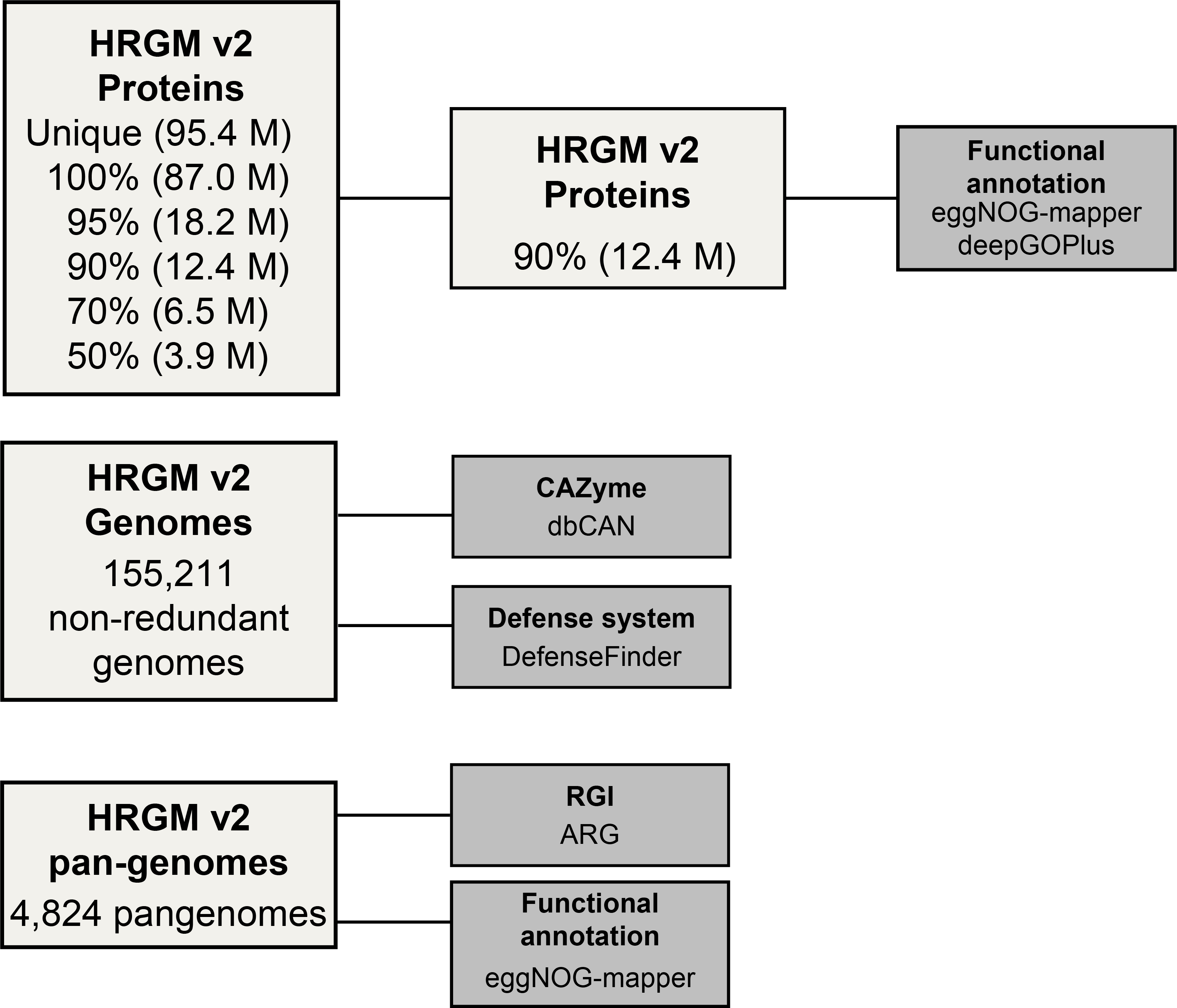
0.HRGMv2_Proteins/
: All protein sequences predicted from all genomes, unique protein sequences after redundancy removal,
and five classes of protein catalogs clustered at different identity thresholds (100%, 95%, 90%, 70%, 50%)
- 0.Redundant_CDS/ : All redundant CDS sequences (549,278,140 coding sequences from 230,632 redundant NC genomes)
- 1.HRGMv2_Unique_Proteins/ : Unique protein sequences after redundancy removal
- 2~6.HRGMv2_{identity}_Proteins/ : Clustered protein catalogs at 100%, 95%, 90%, 70%, and 50% identity thresholds
1.HRGMv2_Pangenomes/
: RGI and eggNOG-mapper results for 4,824 HRGMv2 species (predicted from species-specific pangenomes)
- emapper_results/ : Output of eggNOG-mapper
- rgi_results/ : Output of RGI (Resistance Gene Identifier)
2.HRGMv2_CAZymes/
: Output ofrun_dbcan v4.1.4(standalone version of dbCAN3). CAZyme families were annotated from 155,211 non-redundant genomes.
- For bulk download:download_link_info_cazyme.tsv(full download paths for each non-redundant genome)
3.HRGMv2_Defense_systems/
: Output of DefenseFinder for genome-resolved detection of bacterial defense systems.
- For bulk download:3.HRGMv2_Defense_systems.tar.gz(archive of the full folder)
* Folder structure for 2.HRGMv2_CAZymes/ and 3.HRGMv2_Defense_systems/ follows a 4-level hierarchy to facilitate navigation:
2.HRGMv2_CAZymes/ or 3.HRGMv2_Defense_systems/ ← Root
└── HRGMv2_20XX/ ← Level 1 (group of 100s)
└── HRGMv2_204X/ ← Level 2 (group of 10s)
└── HRGMv2_2040/ ← Level 3 (species-level folder)
├── GENOME008241.tar.gz ← Level 4 (result archive)
└── ...
Present directory - data/protein_catalog
| Name | Last modified | Size | |
|---|---|---|---|
| Parent Directory | - | - | |
| 0.HRGMv2_Proteins | 2025-04-23 20:17:04 | - | |
| 1.HRGMv2_Pangenomes | 2025-04-23 20:38:15 | - | |
| 2.HRGMv2_CAZymes | 2025-04-23 21:07:10 | - | |
| 3.HRGMv2_Defense_systems | 2025-04-23 21:09:41 | - | |
| 3.HRGMv2_Defense_systems.tar.gz | 2025-04-23 21:03:48 | 377 MB | |
| README.txt | 2025-07-21 11:08:26 | 2 KB | |
| download_link_info_cazyme.tsv | 2025-04-23 20:47:16 | 20 MB |